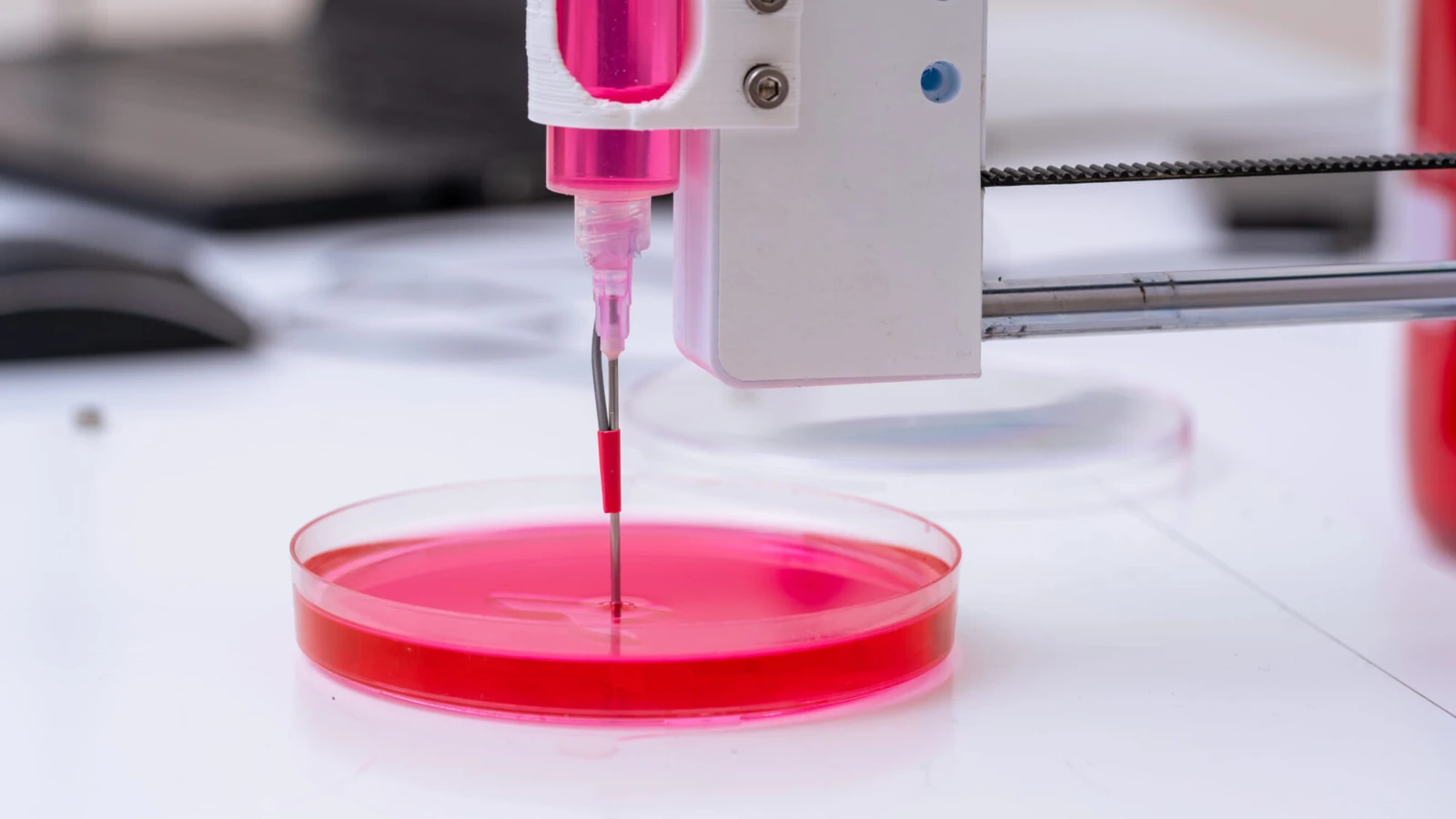

23.02.2023 by Prof. Dr. Michael Gasik (Aalto University Helsinki, Finland)
DMA sobre biomateriales: Ver lo invisible
Un artículo del Prof. Dr. Michael Gasik (Universidad Aalto de Helsinki, Finlandia)
En la actualidad se dispone de muchos tipos de biomateriales para su uso en diferentes implantes, especialmente en casos ortopédicos y dentales. Se utilizan aleaciones metálicas, cerámicas y composites, con o sin células vivas. Existe un creciente campo de aplicación de diferentes andamiajes utilizados en aplicaciones de ingeniería tisular para apoyar y promover la formación de nuevos tejidos, y muchos de ellos se están fabricando mediante (bio)impresión 3D. Se sabe que la regeneración de tejidos biológicos es uno de los retos más exigentes, ya que requiere estructuras de biomateriales con propiedades biomecánicas correctas [1] que imiten el comportamiento in vivo [2]. Unos biomateriales adecuados ayudan al organismo a reconstruir el tejido dañado y minimizan el dolor y el tiempo de cicatrización asociados [3].
Este artículo del Prof. Dr. Michael Gasik (Universidad Aalto de Helsinki, Finlandia) muestra una nueva aplicación de la técnica de análisis dinámico-mecánico (DMA), denominada BEST(Biomaterials Enhanced Simulation Testing), utilizada para caracterizar y mejorar biomateriales y dispositivos médicos; este método va más allá de los análisis viscoelásticos clásicos conocidos.

El profesor Michael Gasik, de la Universidad Aalto de Finlandia (Departamento de Ingeniería Química y Metalúrgica), empezó a trabajar en aplicaciones de análisis térmico en 1985 y lleva casi el mismo tiempo colaborando con NETZSCH-Gerätebau GmbH.
Su trabajo se ha centrado en materiales para aplicaciones de alta temperatura y para la tecnología del hidrógeno. Desde 2000, trabaja activamente con biomateriales, dispositivos médicos y aplicaciones de medicina regenerativa. En 2019, fue nombrado embajador de la Sociedad Europea de Investigación Ortopédica.
El Prof. Michael Gasik es cofundador de Seqvera Ltd. e inventor del método BEST (Biomaterials Enhanced Simulation Testing), que se ha implementado por primera vez en los equipos DMA de NETZSCH.
Un punto central de las actividades de investigación del Prof. Michael Gasik es la determinación de las propiedades mecánicas de los biomateriales. En este contexto, utiliza los datos de DMA generados con un NETZSCH DMA 242 Artemis como base para cálculos posteriores para caracterizar estos materiales. Más información sobre su enfoque:
Retos
Ya se han realizado numerosos estudios y se han recopilado datos clínicos sobre la forma, el diseño y el estado de la superficie de los biomateriales, así como sobre la geometría de los implantes y su adecuación a la calidad y la ubicación de los distintos tejidos. También se notificaron diferencias significativas en materiales implantados aparentemente idénticos pero procedentes de fuentes distintas [4]. La caracterización biomecánica de los tejidos óseos y blandos es más problemática que la de los materiales metálicos, cerámicos y poliméricos. Los conjuntos de datos publicados no suelen basarse en protocolos y condiciones de medición comparables, lo que da lugar a una falta de coherencia. La generalización de estos datos es muy difícil o casi imposible cuando se trata de proporcionar información sencilla, sólida y relevante.
Para la caracterización biomecánica, se suele recurrir a suponer que un material es un tipo de materia elástica o viscoelástica con el fin de aproximar las propiedades a números individuales, normalmente denominados "Módulo elásticoEl módulo complejo (componente elástico), módulo de almacenamiento o G', es la parte "real" del módulo complejo global de la muestra. Este componente elástico indica la respuesta sólida, o en fase, de la muestra que se está midiendo. módulo elástico". Esto, sin embargo, sólo se ajusta a materiales elásticos lineales para deformaciones muy small, y las directrices del NPL [5] enumeran nueve métodos para calcular el Módulo elásticoEl módulo complejo (componente elástico), módulo de almacenamiento o G', es la parte "real" del módulo complejo global de la muestra. Este componente elástico indica la respuesta sólida, o en fase, de la muestra que se está midiendo. módulo elástico que pueden conducir a diferentes valores. Está claro que la gran mayoría de biomateriales y tejidos no son elásticos, por lo que es una simplificación excesiva tratar de reducir artificialmente los datos a unos números fijos: ¿Cuál sería, por ejemplo, el beneficio de conocer el "Módulo elásticoEl módulo complejo (componente elástico), módulo de almacenamiento o G', es la parte "real" del módulo complejo global de la muestra. Este componente elástico indica la respuesta sólida, o en fase, de la muestra que se está midiendo. módulo elástico de la mucosa" que va de 0,1 a 680 MPa según diferentes fuentes?
Por desgracia, las cuestiones relacionadas con los efectos de la inercia (altas frecuencias) o los límites de los instrumentos (bajas frecuencias) no siempre están suficientemente documentadas en los protocolos de ensayo publicados. Incluso si se elimina la inercia del instrumento, la muestra en sí siempre tendrá una inercia finita, que puede producir artefactos por difusión de momento, ondas viscoelásticas y flujos secundarios, todo lo cual puede violar el supuesto de deformación homogénea y lineal [6]. Los modelos más sofisticados tienen un número considerable de parámetros de ajuste artificiales, y existen grandes dificultades experimentales para llevar a cabo dichas pruebas dentro de los estándares, protocolos y métodos de prueba ad hoc existentes [7].
En el caso de procesos como la bioimpresión 3D, hay varios retos que deben superarse, como el control de las propiedades de las biotintas, la gestión del flujo y su efecto en la viabilidad de las células, y la garantía de unas propiedades biofísicas óptimas de las construcciones tras la impresión y en el momento de la implantación [8]. Se requiere una mayor resolución y velocidad con control en el microambiente 3D, y se debe lograr una combinación óptima de propiedades mecánicas y de transporte dentro de la escala espacial y temporal; estas son necesarias en particular para la vascularización limitada por difusión. Las nuevas normativas sobre productos sanitarios (2017/745) exigen que se lleve a cabo una evaluación mecánica adecuada, lo que da lugar a la adhesión a las normativas sobre evaluación de tecnologías sanitarias (2021/2282).
Desafortunadamente, muchos métodos diferentes de pruebas biofísicas arrojan resultados bastante diferentes, y no es fácil obtener propiedades realistas y verdaderas. Las diferencias se deben a muchas razones: contacto desigual, estado de fase, efectos de inercia e inestabilidad elástica, ajuste con modelos asumidos incorrectamente, limitación en la definición de la deformación, falta de una evaluación adecuada del historial de carga, etc. Por lo tanto, es muy importante contar con un enfoque sólido que pueda cuantificar tanto el comportamiento de un biomaterial como su rendimiento en el proceso, en lugar de limitarse a generar algunas cifras concretas.
El concepto BEST
Para hacer frente a estos retos, hemos desarrollado el método patentado BEST(Biomaterials Enhanced Simulation Testing). Puede aplicarse a muchos biomateriales duros y blandos, como hidrogeles, construcciones impresas en 3D y administración controlada de fármacos. Las soluciones BEST se centran en los problemas causados especialmente por ensayos inadecuados y fragmentados, y se establecen sobre un enfoque integrado basado en un principio de causalidad fundamental: "No hubo respuesta por parte de la muestra antes de que se aplicara el estímulo"
BEST se realiza en condiciones controladas con los estímulos coherentes requeridos en el entorno DMA. Evalúa los cambios en las propiedades del espécimen en los dominios del tiempo, la fase y el estímulo [9]. En el postprocesado, BEST integra los datos, convoluciona la historia del espécimen y extrae valores desconocidos, todo ello sin necesidad de que el usuario select el modelo de material (el análisis de datos es esencialmente libre de modelo). Los parámetros invariantes obtenidos con un algoritmo propio de regresión cuántica incorporan la historia del espécimen, mostrando la posición y dirección del desarrollo de un biomaterial [10].
La característica clave de BEST es el procesamiento invariante de los datos DMA, que normalmente permanece inexplorado por el usuario. Este nuevo método supera las limitaciones comunes en la linealidad de las propiedades tisulares en muchos modelos, a saber, una propiedad de escalado (homogeneidad) y una propiedad de superposición (aditividad), que no se dan generalmente para la transformación de Fourier utilizada en la viscoelasticidad lineal.
Por lo tanto, BEST aplica un protocolo de prueba correcto y utiliza métodos idempotentes para extraer parámetros de una sola muestra/prueba, lo que da como resultado datos de alto rendimiento sin el uso de matemáticas complejas (sin necesidad de módulos complejos) ni la suposición de linealidad, y es capaz de reprocesar también otros datos reológicos de forma que no pierdan su valor.
Ejemplo de aplicación de DMA
En el ejemplo que se muestra aquí, el método descrito anteriormente se implementó basándose en mediciones realizadas con un NETZSCH DMA 242 Artemis® para caracterizar las propiedades del hidrogel acrílico para bioimpresión 3D sin utilizar un modelo supuesto. El gel se introdujo en una jeringa de 1 ml con una aguja de 29 G y se colocó en el portamuestras DMA personalizado que se utiliza habitualmente para la flexión; se probó en modo de fluencia escalonada a 25 °C.
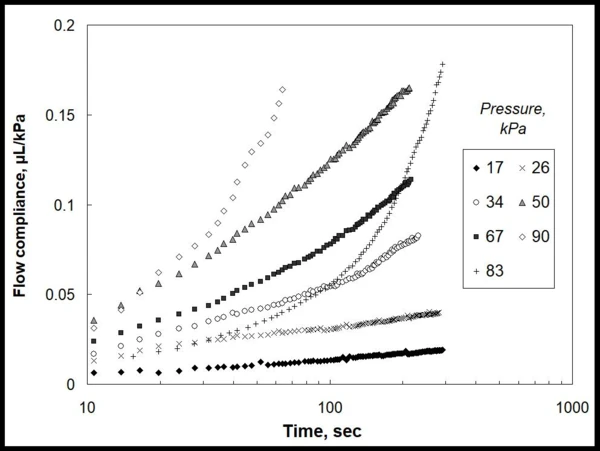
La figura 1 muestra los datos experimentales para la cantidad de gel extruido (µL) a través de una boquilla de aguja definida, normalizada por la presión local aplicada (kPa). Estos datos revelan claramente la no linealidad de la cinética de flujo con el tiempo y la presión aplicada, y no existe una forma directa de select ningún modelo de material que describa estas dependencias.
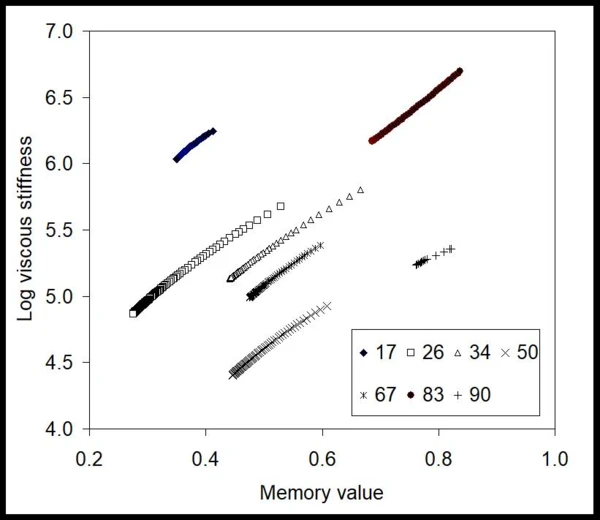
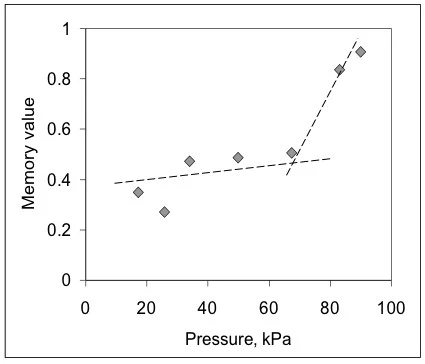
A partir de estos datos, el método BEST extrajo valores invariantes en el tiempo para la rigidez viscosa del gel en estas condiciones de inyección, así como su valor de memoria [9,10] (fig. 2). Aquí, las curvas son casi lineales, y las pendientes de las líneas son casi constantes para todas las presiones aplicadas (números en kPa). Esto significa que el gel, a pesar de mostrar un comportamiento No newtonianoUn fluido no newtoniano es aquel que presenta una viscosidad que varía en función de la velocidad de cizallamiento o de la tensión de cizallamiento aplicada.no newtoniano, es lineal en términos de valores invariantes libres de modelo. También puede observarse que los valores numéricos cambian con la presión aplicada de forma no monótona, lo que revela que puede haber distintos fenómenos limitantes que afectan al flujo. Para ver el efecto del desarrollo del flujo, en la fig. 3 se muestra el gráfico de los valores de memoria frente a la presión aplicada. Este mapa muestra que el gel en la jeringa se enfrenta a fricción, resistencia al flujo y posiblemente efectos de no deslizamiento a bajas presiones, cuando los valores de memoria son muy inferiores a la unidad. Después de unos 65 kPa -el inicio- estos valores aumentan, lo que indica que el gel alcanza un flujo más desarrollado.
El método presentado puede determinar valores invariantes y utilizarlos en la predicción sin modelos de los procesos de bioimpresión 3D, en función de la boquilla, la geometría, la presión, el tiempo y otras condiciones del proceso, sin necesidad de determinar por separado los parámetros reológicos de la tinta. El método BEST genera datos "de primera mano" para el posterior modelado predictivo del proceso de impresión 3D y aplica la misma filosofía para la caracterización de los tejidos y constructos impresos en 3D.
RESUMEN
El enfoque desarrollado demuestra la capacidad de "ver características invisibles" de los materiales y sus interacciones con los estímulos y el entorno. De este modo, el análisis dinámico-mecánico (DMA) puede proporcionar mucha más información que los módulos elásticos y la tangente de pérdida. Mediante el procesamiento BEST, se pueden obtener muchas lecturas para diversos fines (en algunos casos, incluso a partir de una sola probeta o ensayo). Por ejemplo, es posible obtener el módulo de agregación; el tiempo de Deborah característico; la conformidad de fluencia; la difusividad y permeabilidad/permeabilidad efectivas del fluido; el tamaño equivalente del canal para el flujo de fluido en la dinámica; el valor de memoria del material; la presión de hinchamiento; y mucho más en un solo experimento. Y esto va más allá de los biomateriales, ya que la aplicación BEST no requiere modelos ni parámetros de ajuste; además, también puede aplicarse a datos de ensayo ya creados.
Bibliografía:
[1] Hubbell J.A. Nature Biotechnol. 13 (1995) 565-576.
[2] Gasik M. Sci. Techn. Adv. Mater. 18 (2017) 550-562.
[3] Chung C., Burdick J.A. Adv. Drug Delivery Rev. 60 (2008) 243-262.
[4] Gasik M., Lambert F., Bacevic M., Materiales 14 (2021) 2845.
[5] Lord J.D., Morrell R. Measurement Good Practice Guide No. 98; NPL Teddington, Reino Unido (2006)
[6] Ewoldt R.H., Johnston M.T., Caretta L.M. En: Complex Fluids in Biological Systems; Springer, Alemania (2015).
[7] Vrana N.E., Knopf-Marques H., Barthes J. (Eds.) Biomateriales para la regeneración de órganos y tejidos; Woodhead Publ. REINO UNIDO (2020).
[8] Jammalamadaka U., Tappa K. J. Funct. Biomater. 9 (2018) 22
[9] Gasik M., Bilotsky Y. Patente US 10379106 B2 (2019).
[10] Gasik M.Patente US 10809171 B2 (2020).
Contacto:
Prof. Dr. Michael Gasik, Dr. Sci.
Terkko Health Hub, Building 14
Helsinki University Central Hospital Area
Haartmaninkatu 4, FIN-00290 Helsinki
www.seqvera.com
Muchas gracias al Prof. Dr. Michael Gasik por este artículo y la información sobre su trabajo de investigación.







